Large-scale Integrative Taxonomy
Table of Contents
What is LIT?
LIT is a workflow that helps you validate species boundaries by recommending a subset of specimens to check using a secondary data source (e.g. morphology or additional molecular markers) after clustering using COI barcodes. This saves both time and cost while maintaining high taxonomic accuracy.
LIT categorises Molecular Operational Taxonomic Units (mOTUs/clusters) into two types based on:
- Maximum Pairwise Distance (Max P-dist)
- Instability Index
Depending on the cluster type, LIT recommends how many and which specimens to check:
-
Potentially Congruent clusters
Default criteria: Max P-dist < 1.5 AND Instability Index = 1
→ LIT selects the most distant pair of specimens. -
Potentially Incongruent (PI) clusters
Default criteria: Max P-dist ≥ 1.5 OR Instability Index < 1
→ LIT selects the most distant pair and a representative from the largest haplotype.
For more information and detailed explanation about LIT, read Hartop et al. (2022).
LIT Workflow
You should have:
.itvfile from clustering output.htmldendrogram viewer opened
If not, please see the Clustering section under How to use.
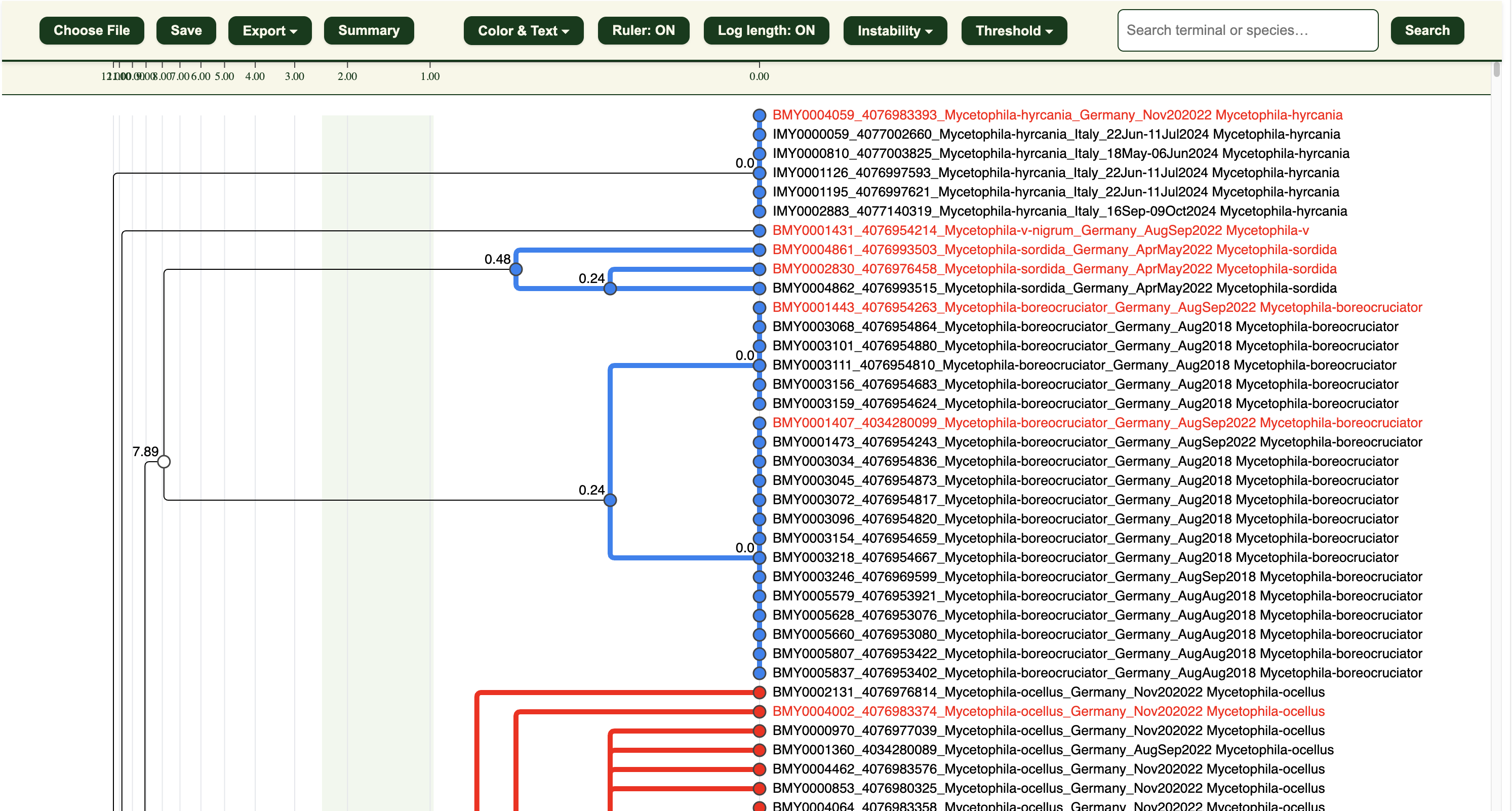
1. Click “Choose file” in the .html dendrogram viewer and upload your .itv file.
You should now see a dendrogram with coloured branches and nodes.
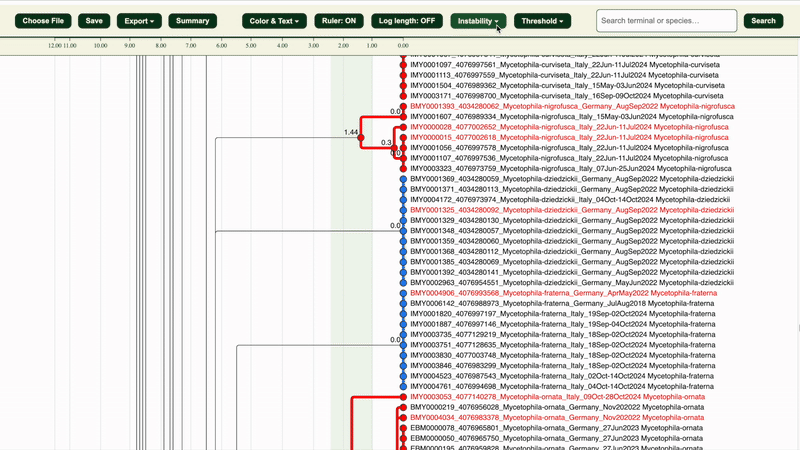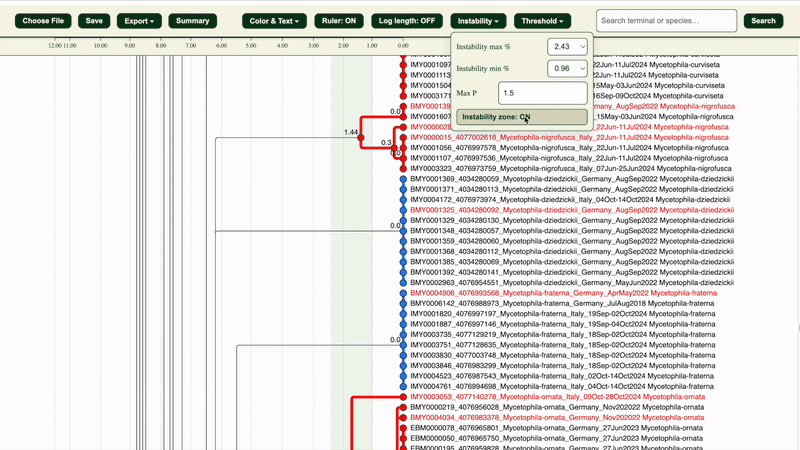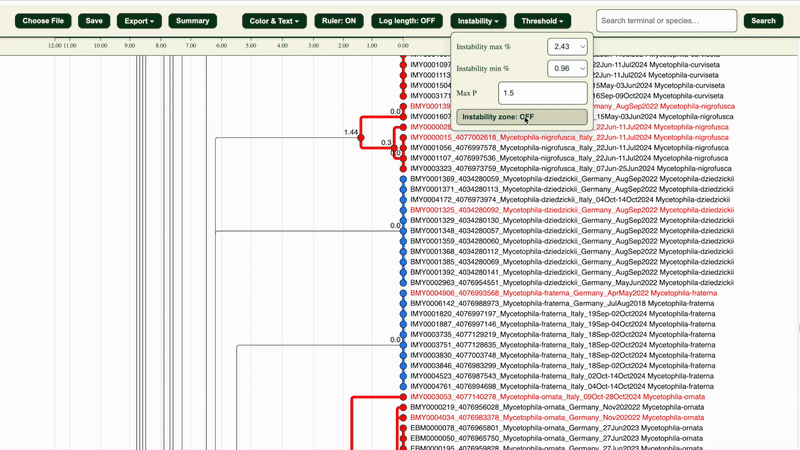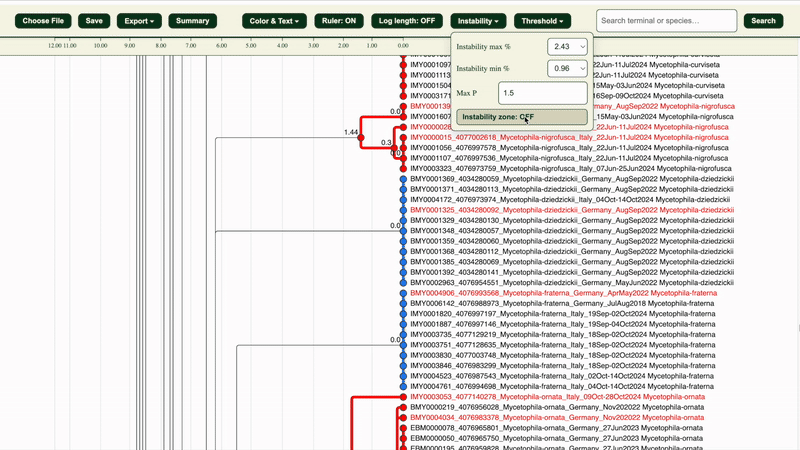
2. Under “Instability” section, adjust the instability maximum and minimum percentages and maximum pairwise distance(Max P) if needed:
- Max P: default is 1.5
- Instability max/min %: defaults are the highest percentages that are less than 3% and 1% respectively (e.g. 2.43% and 0.96%) Both variables affect how clusters are categorised (Potentially congruent or Potentially incongruent).
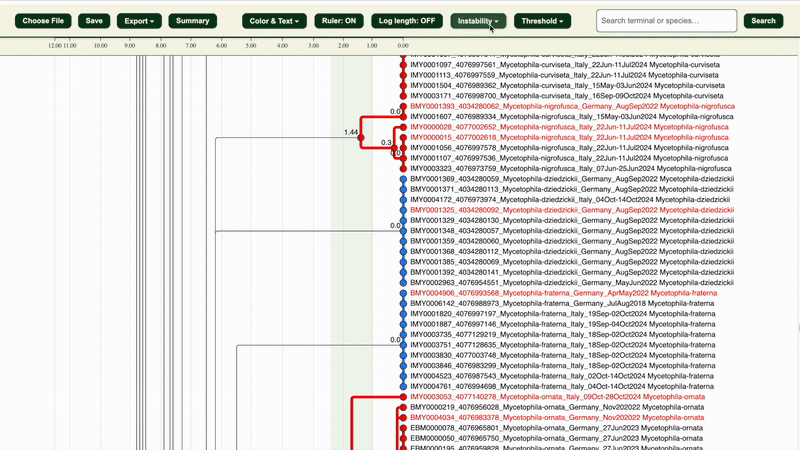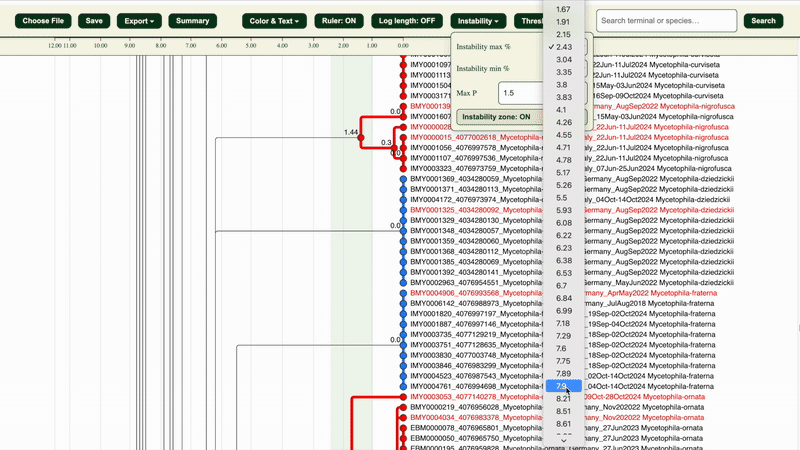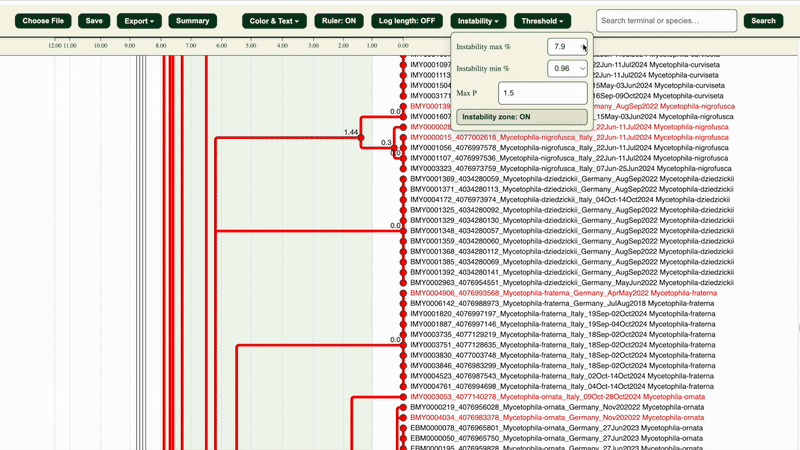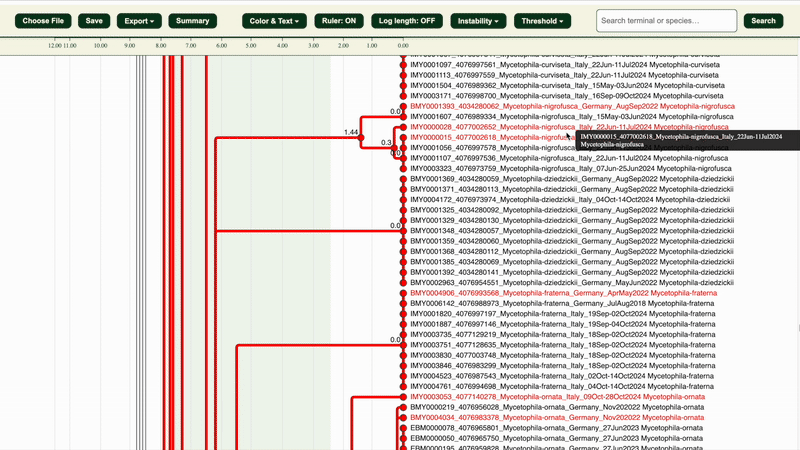
The “Instability zone: ON/OFF” button in the “Instability” section highlights the area within the selected LIT range in green.
3. The dendrogram clusters will be in:
- 🔵 Blue = Potentially Congruent clusters
- 🔴 Red = Potentially Incongruent clusters
The red sequence headers are the LIT-selected specimens for checking.

4. Check the LIT-selected specimens with a secondary data source (e.g. morphology or nuclear markers). Refer to cluster validation case studies below for examples.
If a recommended specimen cannot be checked (e.g. wrong sex, contamination, damage),
right-click the node and assign a reason. Then, select another specimen from the same haplotype. If none are usable, choose one from the next closest haplotype.
For example: If specimen BMY0001528 cannot be verified because it is a female specimen, set the specimen as “Cannot be verified” and “female”. The next most distant haplotype will be BMY0001343. Check that alternative specimen and the other LIT selected specimen, BMY0001521.
5. Track your verification progress by right-clicking the specimen nodes.
You can mark them as verified/cannot be verified, male/female, and other special conditions.

6. Once a cluster is validated as a species, right-click the cluster node
to verify it or assign a species name/code.
✅ The entire cluster (nodes, branches, labels) will turn green.
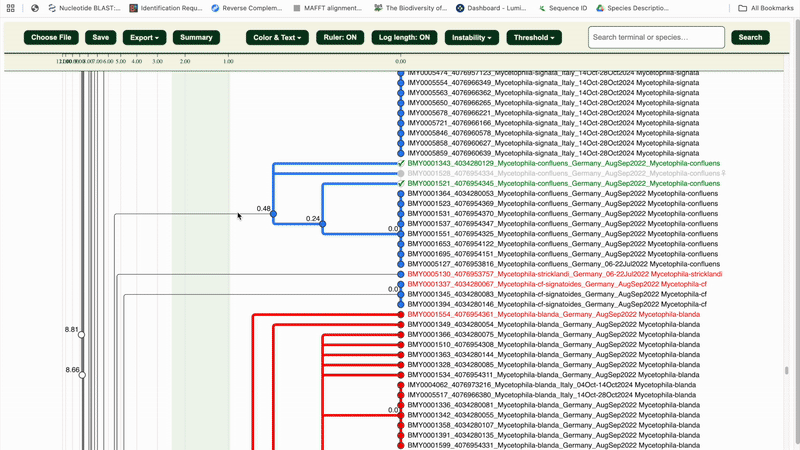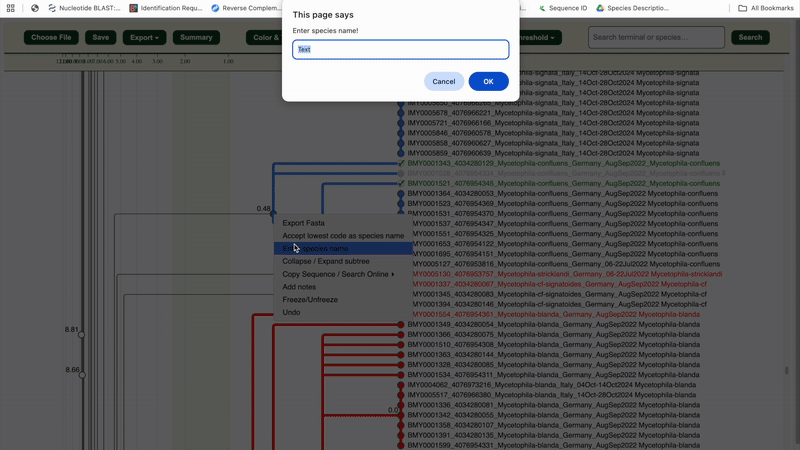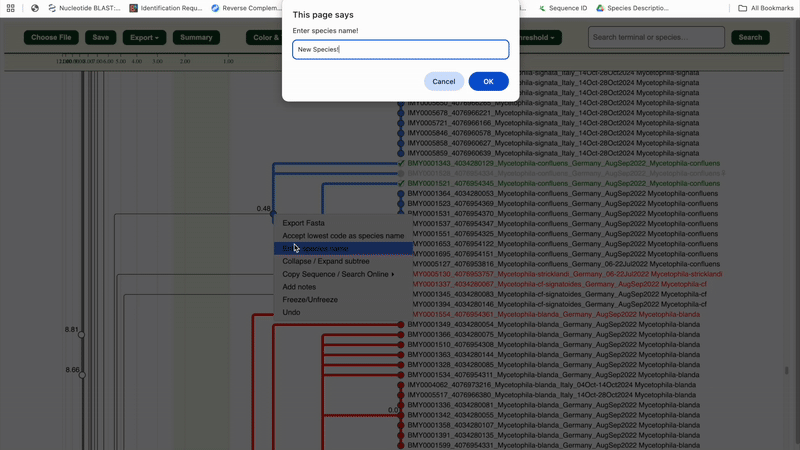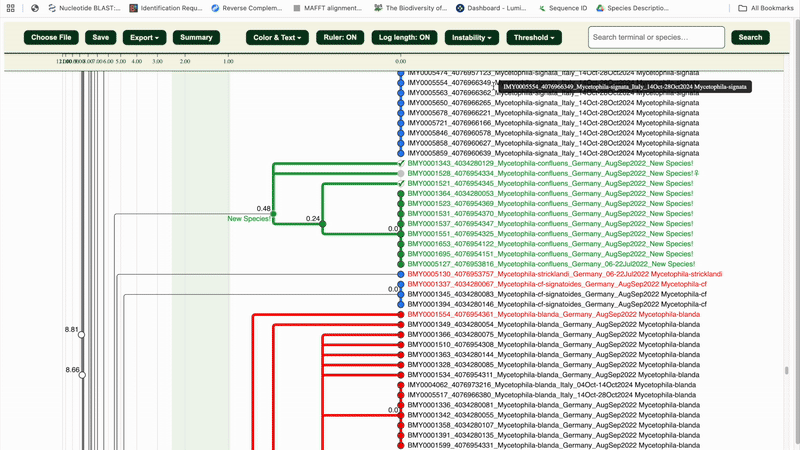
7. Repeat steps 4–6 until your full dataset is reviewed.
8. Want to pause and resume later?
Click the “Save” button to export a .itv file with your progress.
Learn how to save and reload.
9. Export your updated dataset into multiple formats containing information that you have added during the verification process.
LIT cluster validation case studies
The following case studies illustrate how to interpret and validate clusters during the LIT workflow using secondary data sources (e.g. morphology). Each case represents a typical scenario you may encounter.
1. “Blue” Potentially Congruent Cluster
Checking the LIT-selected specimens from a potentially congruent cluster should confirm the cluster as a species. LIT has been tested across multiple Diptera families and other arthropod groups, and we have not observed any cases where a potentially congruent cluster turned out to be incongruent.
For example: IntegraTax suggests the two most distant specimens (Based on pairwise distance) BMY0001528 and BMY0001521 for checking. You will then check the specimens with morphology to verify that both specimens are the same species. Once confirmed, we can verify the entire cluster to be the same species.
2. “Red” Potentially Incongruent Cluster
LIT-selected specimens from a Potentially Incongruent (PI) cluster may represent one or multiple species.
- If your secondary data confirms the selected specimens belong to the same species, you can verify the cluster as a species. For example: In this cluster, IntegraTax suggests 3 specimens for checking (IMY0003053, BMY00004034, and BMY0000188). After verifying that all 3 specimens are the same species, we verified the whole cluster.
- If they represent different species, select additional representatives from each haplotype to verify and split the cluster accordingly. For example: In this cluster, IntegraTax suggests 2 specimens for checking (BMY0000754 and BMY0000951). After checking them, we found that they are separate species. We then proceed to check one representative from every haplotype. This confirmed that the cluster splits at 2.56% into 2 separate species.
3. Lumping Two Clusters into One Species
In some cases, two clusters (regardless of LIT assignment) may fall just beyond the upper LIT threshold (e.g. 3.04% when the threshold is 3%).
If no meaningful differences are found, they can be lumped into a single species. For example: There are 2 3% clusters that are 3.04% apart. After checking the specimens suggested by IntegraTax (BMY0004643 and BNMY0004102), you realised that both clusters are the same species. Right-click the node that encompasses both clusters and enter a species name.
